Warren L. DeLano, Ph.D.
DeLano Scientific
San Carlos, California
USA
warren@delanoscientific.com
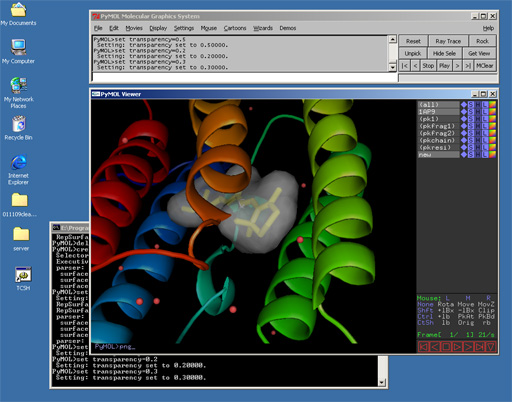
PyMOL is a free cross-platform molecular graphics system made possible through recent advances in hardware 1 , internet 2 , and software development technology 3 .Ā PyMOL provides most of the capabilities and performance of traditional molecular graphics packages written in C or Fortran 4 . However, its integrated Python interpreter endows it with features and expandability unmatched by any traditional package. PyMOL has been released under a completely unrestrictive open-source software license 5 so that all scientists and software developers can freely adopt PyMOL and then distribute derivative works based on it without cost or limitation.

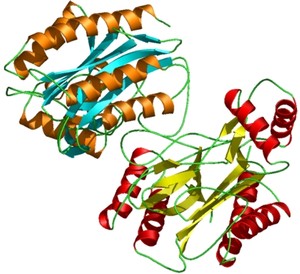
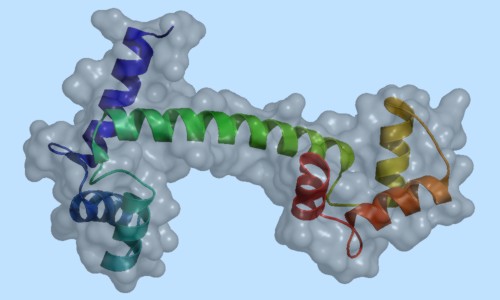
PyMOL supports most of the common representations for macromolecular structures: wire bonds, cylinders, spheres, ball-and-stick, dot surfaces, solid surfaces, wire mesh surfaces, backbone ribbons, and cartoon ribbons which are comparable to those generated by Molscript 6.
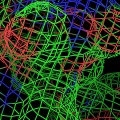

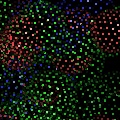
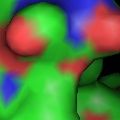



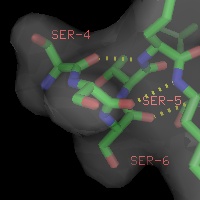
Labels can be displayed for atoms, and dashed bonds can be used to indicate hydrogen bonding interactions and distances. Surfaces can be transparent, and molecules can be loaded from PDB files as well as several other common file formats.
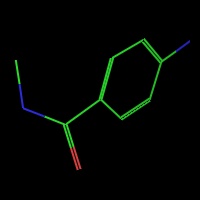
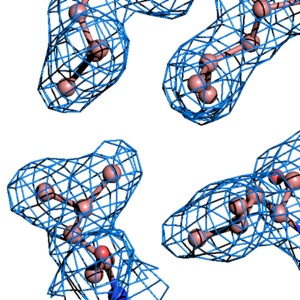
PyMOL reads CCP4 and X-PLOR 7 map files and can display multiple arbitrary bricks of electron density within each map.Ā PyMOL also has the ability to "carve" out electron density around any selection of atoms to create figures which show only localized electron density.
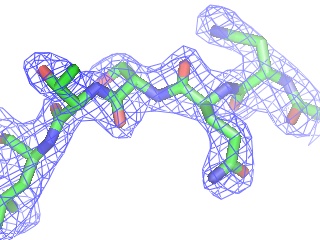
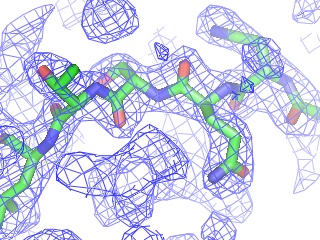
Provided that a structure has been loaded from a PDB file with correct unit cell and space group parameters 8 , PyMOL can generate symmetry-related molecules. This was made possible from R.W. Grosse-Kunstleve's generous contribution of the SgLite package 9 .Ā At present, symmetry-related molecules are treated as independent objects, not as virtual images of the original object.
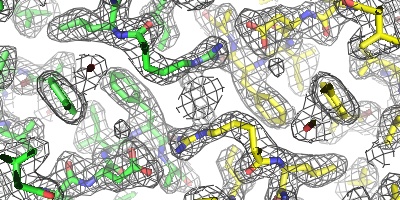
PyMOL contains a rudimentary electron density "wizard" (written in Python) which can be used to quickly navigate through one or more electron density maps surrounding an atomic model. This wizard enables one to move and regenerate multiple meshes merely by CTRL-middle-clicking on the atom to center.
A powerful time-saving feature found in PyMOL is its integrated ray-tracing engine.Ā With a single mouse-click, any view displayed in the program can be immediately converted into a publication quality figure, complete with lighting, specular reflections, and shadows.Ā This obviates the need for time-consuming efforts involving command-line programs such as Molscript 6 , Raster3D 4 , and ImageMagick 10 . With PyMOL, what you construct in real-time 3D is exactly what you will get when you ray-trace (except for labels), and you can import the resulting images directly into Microsoft PowerPoint.
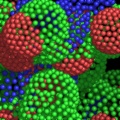
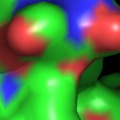
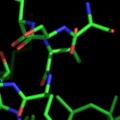
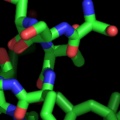
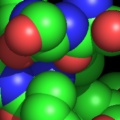
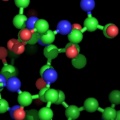


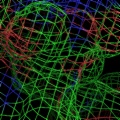


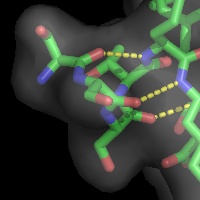
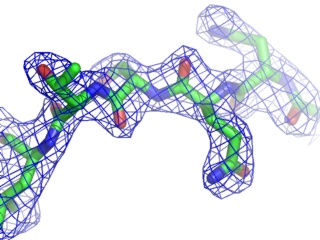
Although PyMOL's built-in ray-tracer is quite good, PovRay 11 support has recently been added, and it can now be used as a replacement renderer for generation of the highest quality images.Ā Both OpenGL and ray-traced images can be output from PyMOL using standard Portable Network Graphics (PNG) files.
PyMOL's native cartoon ribbons are very similar to Molscript's, but if genuine Molscript output is required, PyMOL can read Molscript output in Raster3D format to perform rendering. The advantage of using PyMOL instead of Raster3D is that it allows users to orient and combine multiple objects in real-time 3D prior to rendering.ĀThe quality of the resulting images is comparable using either approach.
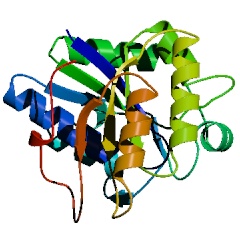 Molscript cartoon ribbons rendered with Raster3D |
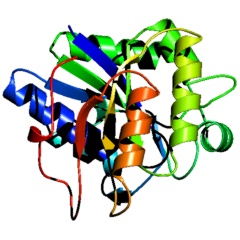 Molscript cartoon ribbons rendered with PyMOL |
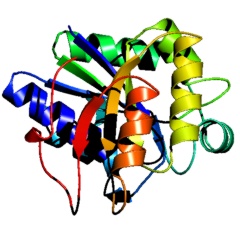 PyMOL cartoon ribbons rendered with PyMOL |
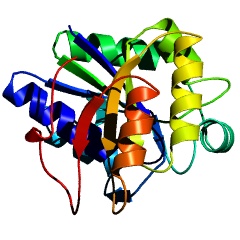 PyMOL cartoon ribbons rendered with PovRay |
PyMOL supports multiple atom selection syntaxes which permit concise atom specifications: "C/143/CA", "C/PHE/", or "*/CA", as well as extended algebraic expressions resembling those found in X-PLOR and CNS 12:Ā"(byres ((resi 125 and chain A) around 5))".Ā For most operations in PyMOL (such as coloring, zooming, or changing representations), atom selections can be used interchangeably with molecular objects.Ā Selections can also be defined using the mouse in several different ways, including using a rectangular "lasso" around visible atoms.Ā All atom selections can be visualized directly in the 3D window, which makes them easy to understand and verify.
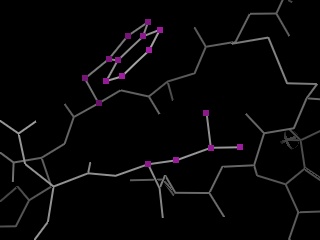
PyMOL supports molecular editing at several different levels.Ā At the simplest level, it allows the user to create new objects out of atom selections which may span any number of other objects.Ā Thus, removing atoms from an object or combining separate objects is trivial.Ā Bonds can then be formed between atoms in the combined object.
PyMOL also supports conformational editing within objects using an intuitive click-and-drag interface (when in "editing" mode).Ā For example, to rotate about a given bond torsion, one merely has to CTRL-right-click on a bond and then CTRL-left-click to drag atoms on either side of the bond.Ā Bonds, angles, torsions, and positions are all editable using this kind of click-and-drag operation.
PyMOL also enables rudimentary chemical editing of structures on an atomic basis using mouse clicks and CTRL-key combinations to delete, replace, or grow new molecular structures.Ā For example, to make tyrosine out of phenylalanine, one would simply CTRL-middle-click on the para-hydrogen and type CTRL-O to replace it with a hydroxyl.
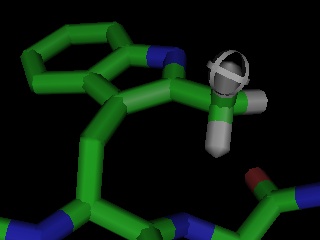
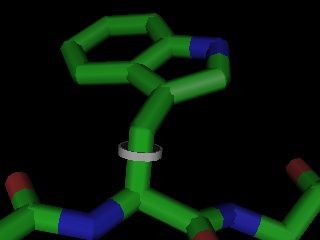

Given PyMOL's existing editing capabilities, a full featured molecular modeling tool could now be implemented on top of PyMOL using only the Python language, without any C or C++ coding.Ā The Python based mutagenesis "wizard" included with PyMOL proves this concept and shows one way such a builder might operate.Ā A crystallography-oriented model building tool is a top development priority, and PyMOL should become suitable for this task in mid-2002. Some ambitious users are already using PyMOL to carry out limited conformational changes late in refinement.
PyMOL was designed from the ground-up to accommodate multiple atomic coordinates for each atom.Ā Thus, it is possible to load trajectories and conformational ensembles directly into PyMOL for dynamic visualization.Ā These can be viewed and rendered using all of the built-in representations, and there is even a facility present for constructing movies as a programmable sequence of molecular states.Ā
PyMOL can then output images of these states as a series of numbered files for assembly into QuickTime or AVI movies.Ā Ray-traced animations can also be previewed with PyMOL by rendering all frames into memory and then paging through them at machine speed.Ā
PyMOL does not require a graphical user interface in order to run.Ā With a simple command line switch, the program can be launched in command-line mode, which resembles a Python interpreter possessing all of PyMOL's built-in rendering and editing functions.Ā One example use of this feature would be to farm out rendering of a molecular animation to a cluster of Linux workstations for parallel processing.
PyMOL support two related control languages: Python and the PyMOL command language. The PyMOL command language is merely a series of Python function calls with implicit quoted arguments and implicit parentheses. This syntax makes the power of Python directly accessible to non-programmers, and it provides a familiar feel for experienced users of crystallographic software. Because the PyMOL scripting language falls back on Python for evaluation, it can be thought of as a superset of the Python language.
Although documentation on PyMOL's features is currently incomplete, a 129+ page user's manual can be downloaded from the PyMOL web site and will help in getting started.Ā PyMOL also supports online help within the program and contains command and argument auto-completion and inference logic to make command-line usage tractable even for novices.
By default, PyMOL starts up like a typical stand-alone molecular graphics program. However, command line options can be used to change this behavior.Ā While PyMOL ships with a simple Tcl/Tk-based "external" graphical user interface (GUI) and a primitive OpenGL-based "internal" GUI, all such features can be disabled and replaced with new components from an external package. Other applications can simply utilize PyMOL as a molecular display window and provide their own external menus, windows, dialog boxes, and controls. These can be constructed using toolkits such as Tkinter (Tcl/Tk) 13 , wxPython (wxWindows) 14 , Qt 15 , MS-Windows 16 , or Mac OSX 17.
PyMOL also allows developers to add additional geometries into the 3D viewing environment using either PyOpenGL 18 (via Callback Objects) or PyMOL's Complied Graphics Objects (CGOs), which represent streams of OpenGL-like drawing commands.Ā The advantage of CGOs is that geometries specified using them can automatically be rendered efficiently in OpenGL using lines and triangles and simultaneously conveyed to the ray-tracer as analytical spheres and cylinders.Ā PyMOL uses CGOs internally for displaying cartoon ribbons and for emulating Raster3D.
So long as external applications are developed using PyMOL's Python-based application programming interface (API), it should be possible for packages to maintain compatibility even as each package evolves independently.Ā PyMOL is thus an attractive platform for any kind of molecular software development project which requires molecular visualization.
PyMOL is an independent software development project of DeLano Scientific 19 , a sole proprietorship based in San Carlos, California, USA.Ā Because PyMOL has been given away for free, DeLano Scientific has a microscopic budget, and nearly all PyMOL development has been performed using home equipment and uncompensated personal time. Unlike comparable academic open-source software development efforts, DeLano Scientific receives no direct support from university or government sources for PyMOL.
Instead, funding for PyMOL development comes directly from the voluntary contributions of PyMOL users and developers who are asked to directly support PyMOL out of their own self-interest. Nearly all biomolecular scientists will benefit from the existence of a powerful and universally available graphics program, so everyone has a reason to contribute. While donations or participation are not legally or ethically required to use PyMOL, they encourage development and are what will make future versions possible.
The outlook for PyMOL is very bright now that robust molecular graphics capabilities are present within the program, and since hundreds of scientists, if not thousands, have already started using PyMOL in their research 20 .Ā Even though PyMOL's C source code was developed extremely rapidly and does not meet professional coding standards, most of the functionality in PyMOL is exposed through a documented Python API. This breakthrough for molecular graphics enables facile reuse of PyMOL source code in many different contexts, without requiring an understanding of it.
Future efforts will continue on a number of fronts, from improving documentation, to adding important new features and polishing the programming interface.Ā There are PyMOL-related efforts ongoing in crystallography, computational chemistry, molecular modeling, simulation, genomics (threading/homology modeling), and education. Eventually PyMOL will become part of an open full-featured molecular computing environment 21.
PyMOL can be downloaded for free via the internet at http://www.pymol.org . The web site contains a variety of other useful information, such as a copy of the manual, information about the PyMOL mailing list, recent news, and related links.Ā PyMOL currently runs on a variety of common platforms: Windows, Linux, IRIX, Mac OSX, and Tru64 Unix. Binaries are usually available for Windows and Linux.
1. ĀThanks to nVidia's fast and low-cost 3D graphics chips, which support Windows, Macintosh, and Linux http://www.nvidia.com. Graphics cards with nVidia chips can cost less than $100 but outperform common Silicon Graphics hardware for crystallography tasks. Complete molecular graphics workstations can now be built for about $600 (or for about $1000 if one needs stereo graphics).
2. Thanks to the SourceForge open-source development infrastructure, based around the concurrent versioning system (CVS).Ā SourceForge reduces software distribution costs to near zero, and has greatly assisted open-source software development http://www.sourceforge.net .
3. Thanks to the languages Python http://www.python.org, C, and OpenGL http://www.opengl.org, and to various software development strategies inspired by the Extreme Programming (XP) approach: low complexity, minimalist implementation, continuous integration, rigorous testing, and routine user participation http://www.extremeprogramming.org.
4. Examples include: Grasp http://trantor.bioc.columbia.edu/grasp, MidasPlus http://www.cgl.ucsf.edu/Outreach/midasplus/index.html, O http://xray.bmc.uu.se/~alwyn , Molscript http://www.avatar.se , Raster3D http://www.bmsc.washington.edu/raster3d/raster3d.html, and MacroModel http://www.schrodinger.com/Products/macromodel.html .
5. PyMOL is released under the "Python" version 1.5.2 license.Ā In essence, PyMOL source code can be used in any way and in any context so long as DeLano Scientific's copyright notices are not removed.Ā See the PyMOL distribution for license details.
6. Molscript, a commercial package available at http://www.avatar.se, has set the standard for molecular graphics for many years. PyMOL comes very close to reproducing Molscript-quality cartoon ribbons in a free and open-source context.Ā Ā
7.
X-PLOR home page http://atb.csb.yale.edu/xplor8. Information on the PDB file format can be found at http://www.rcsb.org/pdb/info.html.
9. An updated version of SgLite is now available as part of the computational crystallography toolbox (cctbx) http://cctbx.sourceforge.net.
10. Imagemagick is an essential image conversion and display tool http://www.imagemagick.org.
11. Persistence of Vision home page http://www.povray.org
12. The Crystallography and NMR System home page http://cns.csb.yale.edu
13. Tkinter http://www.python.org/topics/tkinter is a Python implementation of the Tcl/Tk API http://tcl.activestate.com
14. wxPython http://www.wxpython.org is a Python implementation of the wxWindows API http://www.wxwindows.org.
15. Qt is a cross-platform Windowing library subject to certain commercial restrictions http://www.trolltech.com.
16. The Microsoft Windows environment http://www.microsoft.com.
17. The Mac OSX environment http://www.apple.com/macosx
18. PyOpenGL is a Python implementation of the OpenGL API http://sf.net/projects/pyopengl.
19. Please note the important PyMOL legal disclaimer: WARREN LYFORD DELANO AND DELANO SCIENTIFIC DISCLAIM ALL WARRANTIES WITH REGARD TO THIS SOFTWARE, INCLUDING ALL IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS. IN NO EVENT SHALL WARREN LYFORD DELANO OR DELANO SCIENTIFIC BE LIABLE FOR ANY SPECIAL, INDIRECT OR CONSEQUENTIAL DAMAGES OR ANY DAMAGES WHATSOEVER RESULTING FROM LOSS OF USE, DATA OR PROFITS, WHETHER IN AN ACTION OF CONTRACT, NEGLIGENCE OR OTHER TORTIOUS ACTION, ARISING OUT OF OR IN CONNECTION WITH THE USE OR PERFORMANCE OF THIS SOFTWARE.
20. According to SourceForge statistics in late November, 2001, PyMOL and related files had been downloaded over 12,000 times and generated over 84,000 web site hits since April, 2000.
21. The FreeMOL project aims to create an open alternative to commercial packages like Sybyl (Accelrys), Insight II (Accelrys), Cerius2 (Accelrys), and the Molecular Operating Environment (Chemical Computing Group) http://www.bioinformatics.org/freemol .